About Us
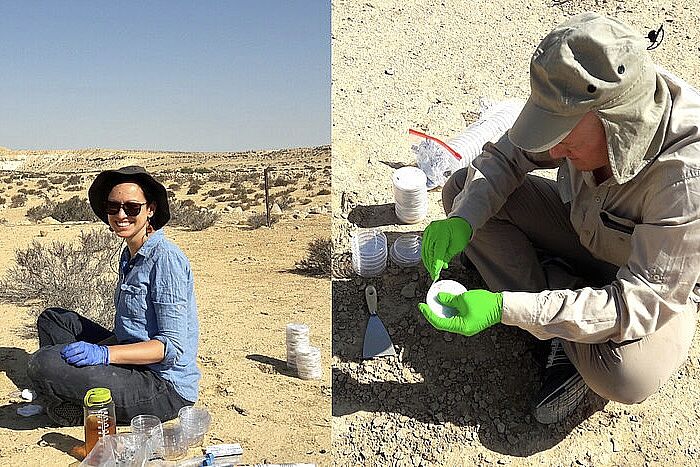
The Division of Microbial Ecology (DOME) at the Centre for Microbiology and Environmental Systems Science (CeMESS) is a team of microbiologists and molecular biologists. Our mission is to unravel the complexities of microbial life and its profound impact on our health and environment.
Our research explores the evolution, ecophysiology, interactions and functions of microbes across a broad range of systems. We apply cutting-edge methods, from 'omics analysis, to single-cell isotope probing, to microbial imaging, to computational community modelling.
Research
At DOME, we are committed to nurturing the next generation of scientists at every level of education, including undergraduate and doctoral programs, as well as intensive methods-based courses.